TLDR
- Vocal Command Processing for Automated Cell Culture Systems
- Cell Painting Analysis: High Content Morphological Profiling of Gene X Impact on Prostate Cancer Lines
- Statistics in Language: Large-Scale Corpus Analysis
- Ventral-Dorsal Stream Interactions in Functional Object Grasps
- Night at the Museum
- Epigenetics of Echolocation
- Decentralized Systems and Swarm Behavior
Vocal Command Processing for Automated Cell Culture Systems
Developed a client-server architecture for offline vocal command processing for automated cell culture systems. Implemented wake word detection using Whisper (tiny model) and command transcription using Whisper (base model) for speech recognition. Built command comprehension pipeline using local LLM to extract structured data (experiment type, plate coordinates, timing parameters) from natural language commands and convert them to JSON format for the automation scheduler.
Designed multi-threaded client application with continuous background listening for wake word activation and on-demand command processing. Maintained and extended multi-threaded automation software for task management and SQL database integration (Tecan), enabling vocal control to start, stop, pause schedules, or add experiments to the automation queue.
Cell Painting Analysis: High Content Morphological Profiling of Gene X Impact on Prostate Cancer Lines
Conducted a novel high-throughput screening image-based Cell Painting assay on BPH1 (Human Benign Prostatic Hyperplasia) and LNCaP (Lymph Node Carcinoma of the Prostate) Gene X knockout and wildtype cell lines to investigate the role of Gene X in prostate cancer progression. Used 6 fluorescent dyes in 5 channels to identify around 700 morphological features of cells pertaining to size, shape, stain intensity, and textural pattern of each of the 8 main organelles.
Created several large datasets using a variety of cell segmentation techniques and trained a Random Forest Classifier model to predict whether a given cell is a wildtype or knockout for the gene, provided the respective Cell Painting data. After extensive image and data analysis using standard statistical techniques in addition to machine learning methods, successfully established a distinct morphological profile of Gene X.
Statistics in Language: Large-Scale Corpus Analysis
Investigated the role of local versus global co-occurrence statistics in semantic knowledge acquisition from child-directed speech. Processed large-scale language corpora (~2.8B words) to analyze co-occurrence structure in natural language input, testing the hypothesis that global co-occurrences (word pairs in similar linguistic contexts) are as exploitable as local co-occurrences (word pairs appearing close together) for semantic learning.
Implemented Pointwise Mutual Information (PMI) and Latent Semantic Analysis (LSA) to model semantic similarity and quantify the semantic information encoded by both local and global statistics. Built NLP pipelines using NLTK and spaCy for corpus parsing, tokenization, and syntactic analysis. Findings demonstrated that global co-occurrences are frequent in child-directed speech and encode patterns of similarity that could support structured semantic representations, challenging accounts that prioritize only local statistics.
Co-authored poster and abstract accepted to CogSci2025: Are Global Statistics Discarded Statistics?
Ventral-Dorsal Stream Interactions in Functional Object Grasps
Investigated how the brain stores and manipulates object knowledge through behavioral training and neuroimaging. Designed experiments testing whether participants could learn to identify balance points of objects with color-weight correlations through repetitive grasping tasks, and whether this learning would generalize to novel objects with similar features.
Conducted behavioral studies where participants learned to pick up foreign objects with various colors and weights, then tested their ability to identify balance points before and after training using both physical and virtual tasks. Processed high-resolution fMRI data to examine neural mechanisms underlying object knowledge acquisition and the interaction between ventral (object recognition) and dorsal (action planning) visual streams. Poster accepted at Cognitive Neuroscience Society in Toronto (April 2024): Ventral-Dorsal Stream Interactions Supporting Functional Object Grasps.
Night at the Museum
Created original game using Python with original graphics and algorithms. Implemented AI for maze generation, backtracking, and pathfinding.
Epigenetics of Echolocation
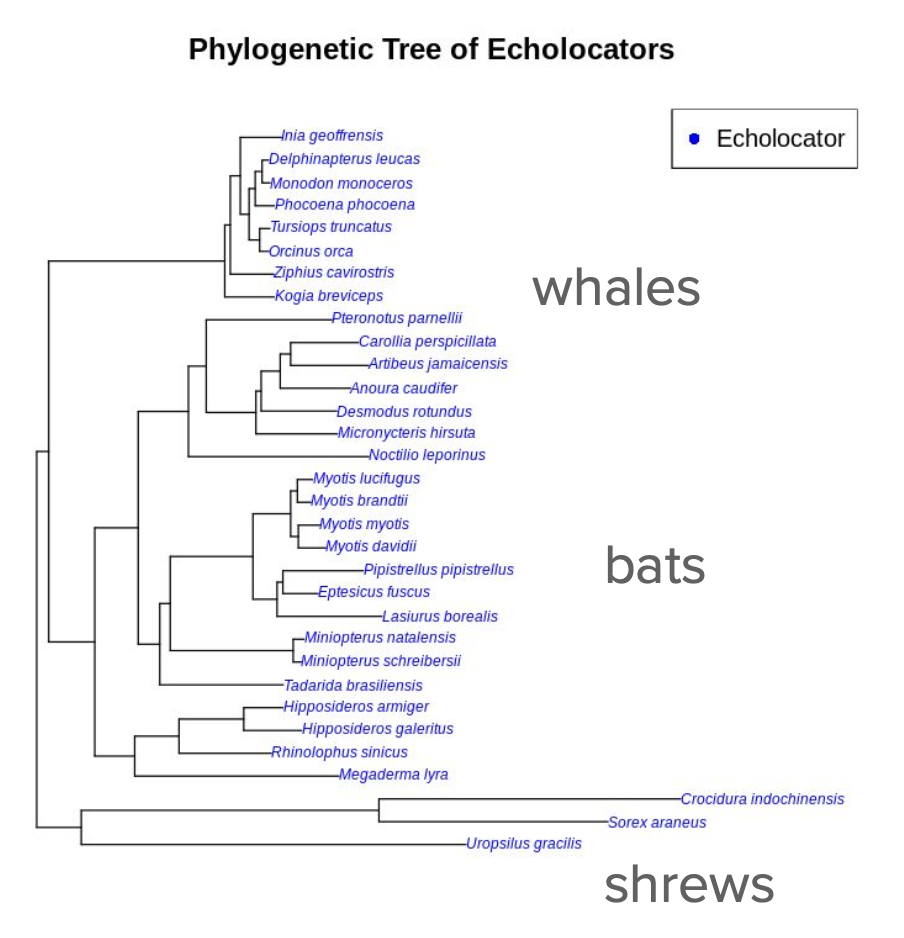
Investigated epigenetic mechanisms underlying echolocation across 32 mammalian species (bats, whales, shrews) using open chromatin analysis in oligodendrocytes and parvalbumin neurons. Developed computational pipeline to identify echolocation-associated chromatin peaks: calculated differences between echolocating vs. non-echolocating species, performed phylogenetic linear modeling (phylolm) to assess correlations, and identified top positively/negatively associated peaks.
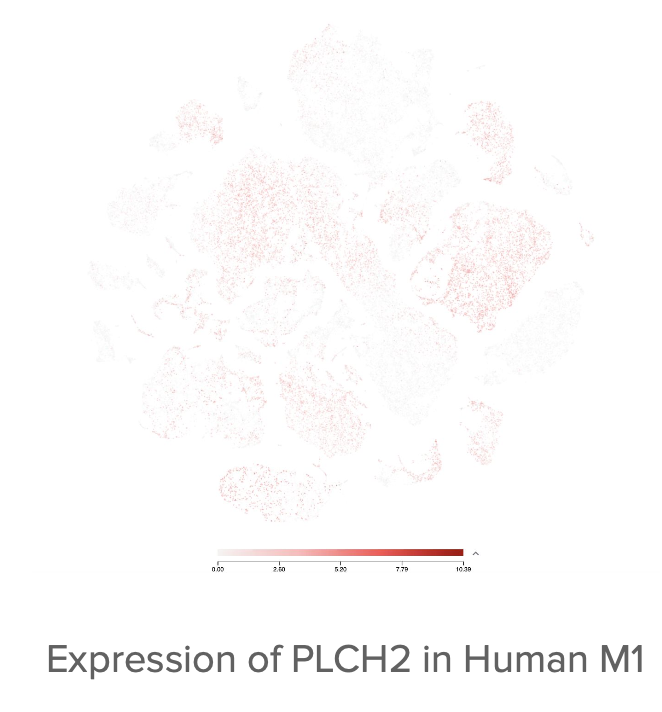
Conducted gene ontology (GREAT) analysis on top 200 positive and negative peaks for both cell types, revealing significant associations with muscle relaxation processes (oligodendrocytes) and cell junction functions (parvalbumin neurons). Identified key genes of interest: AGGF1 (angiogenic factor, muscle development), PLCH2 and PANK4 (neural processing, vision impairment), MIR143/MIR145 (muscle cell differentiation), and PPP1R2B (signal transduction).
Visualized gene expression patterns using UCSC Browser and Allen Brain Atlas, linking epigenetic changes to echolocation through vocal muscle development, enhanced neural processing, and vision inhibition.
Decentralized Systems and Swarm Behavior
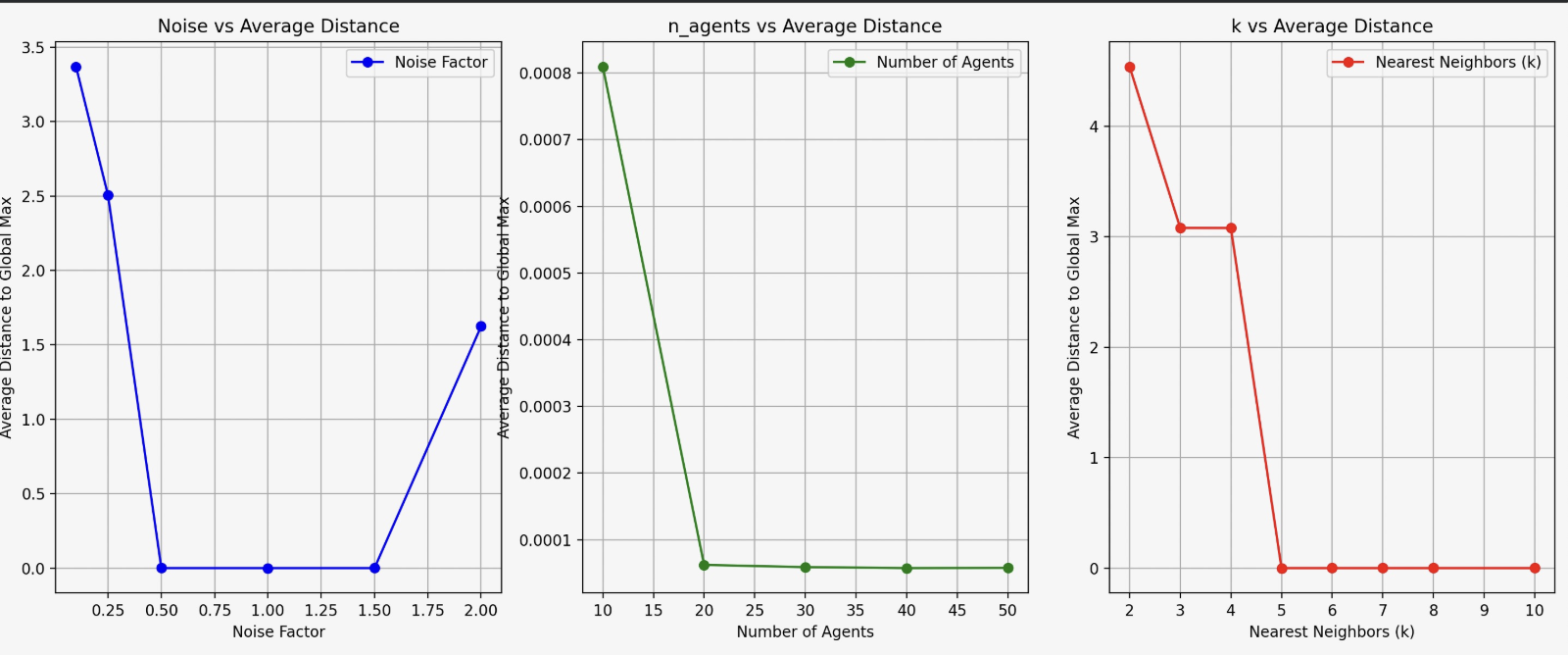
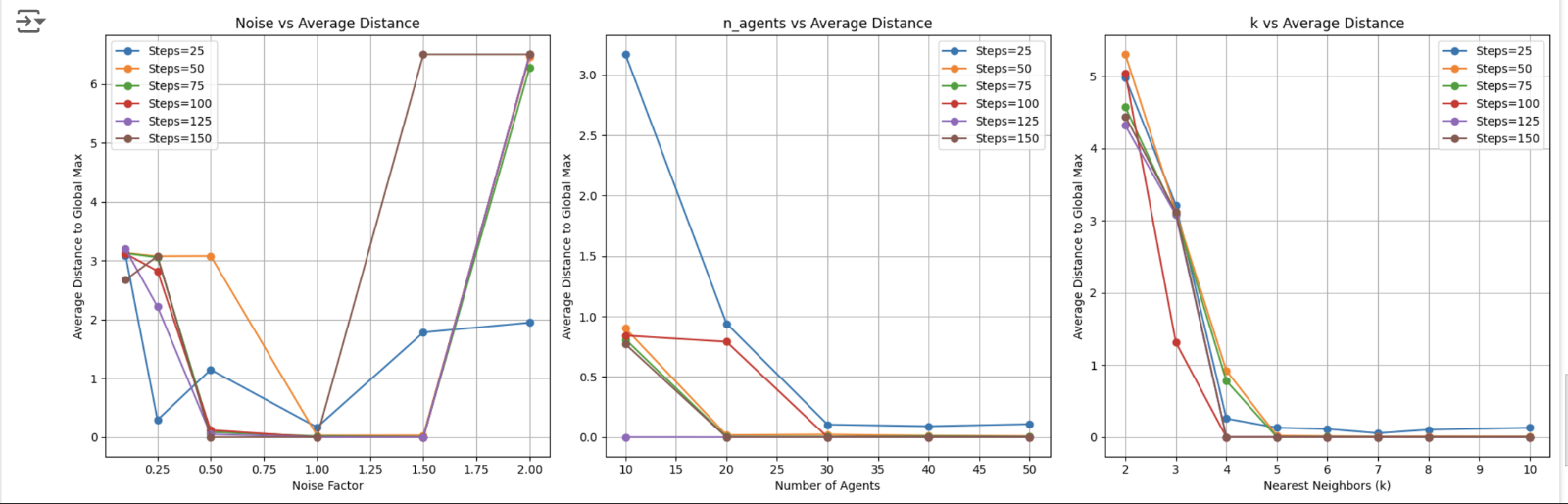
Modeled decentralized systems inspired by natural flocking behaviors (birds, fish schools) to study exploration efficiency in 2D landscapes with multiple local maxima. Implemented agent-based simulation with local communication using KDTree for k-nearest neighbor interactions, uncertainty-based noise computation, and adaptive position updates toward believed global maximum.
Tested three hypotheses: optimal agent count for exploration efficiency, moderate noise levels balancing certainty and diversity, and optimal neighbor count (k) balancing information sharing and decision clarity. Optimized parameters (noise factor, number of agents, k neighbors) by measuring average distance to known global maximum across multiple trials, using 75th percentile to avoid outlier effects.
Key findings: moderate noise (0.5-1.0) optimal for escaping local maxima; optimal agent count ~30 with diminishing returns; k=4 neighbors optimal for information sharing; all three hypotheses validated. Results demonstrate how noise, swarm size, and local interactions balance individual behaviors with collective success, with applications to robotics, AI, and neuroscience.
Created visualization system with contour plots of objective function landscapes and animations of agent movements over time.